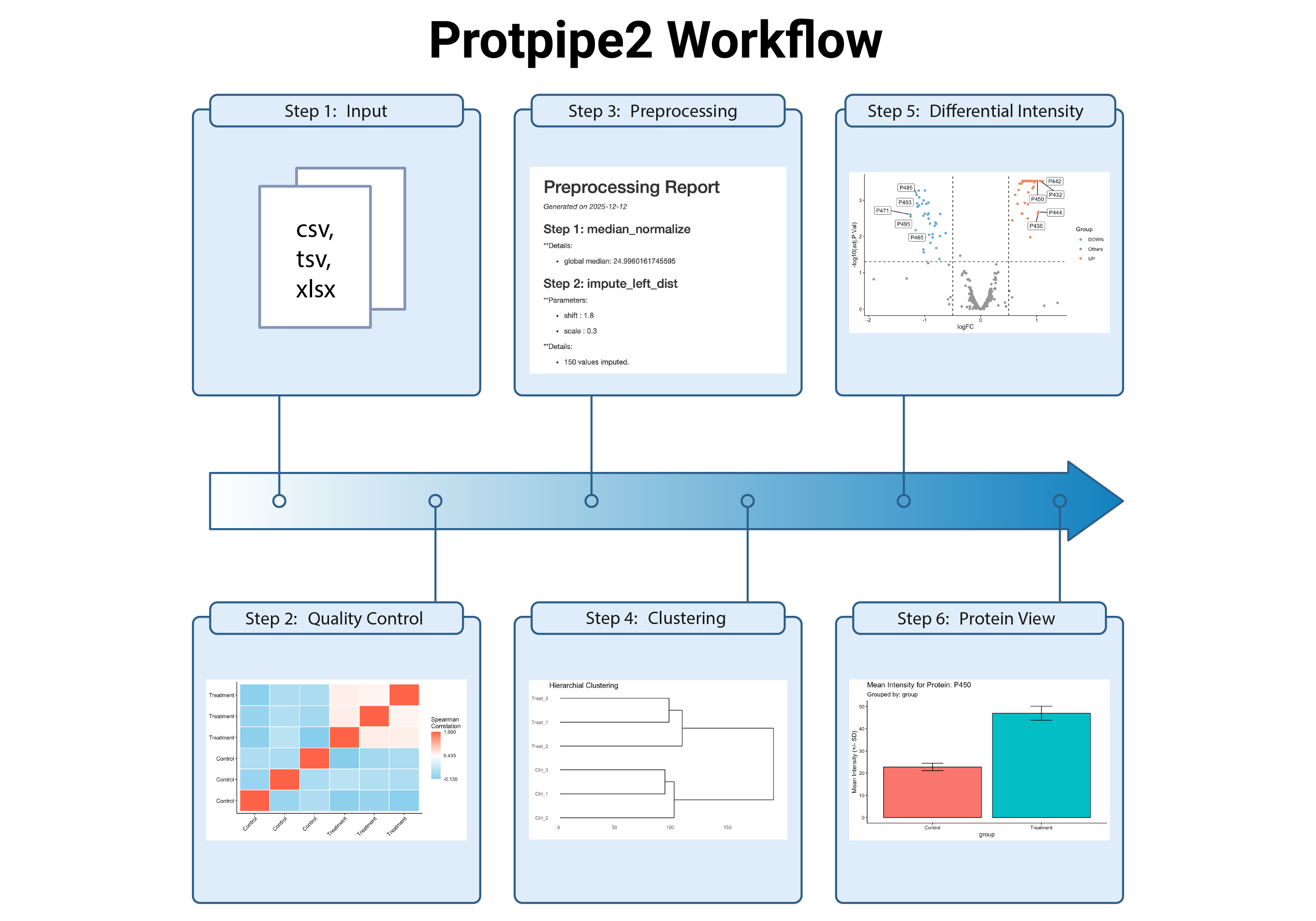
ProtPipe2 provides reproducible workflows for downstream proteomics analysis through the ProtPipe R package and a Shiny interface. The package supports data import into SummarizedExperiment, quality control, filtering, normalization, imputation, batch correction, dimensionality reduction, clustering, differential intensity analysis, and abundance profiling. It also includes visualization and pathway or gene set enrichment utilities for interpretation of protein-level results.
Installation
if (!requireNamespace("pak", quietly = TRUE)) {
install.packages("pak")
}
pak::pak("NIH-CARD/ProtPipe2")
library(ProtPipe)The package installs a minimal core dependency set by default. Some optional features require extra packages:
-
ProtPipe::run_protpipe_shiny()requires the Shiny app dependencies. - Pathway enrichment functions require Bioconductor annotation and enrichment packages.
- SomaScan, Olink, and Excel import paths require their corresponding reader packages.
If one of those optional dependencies is missing, ProtPipe now stops with a direct message naming the package to install.
If the installation step fails on a Bioconductor dependency, install BiocManager and retry after installing the missing package:
install.packages("BiocManager")
BiocManager::install("<missing-package>")